NHITS’s inputs are static exogenous , historic exogenous , exogenous available at the time of the prediction and autoregressive features , each of these inputs is further decomposed into categorical and continuous. The network uses a multi-quantile regression to model the following conditional probability: In this notebook we show how to fit NeuralForecast methods for binary sequences regression. We will: - Installing NeuralForecast. - Loading binary sequence data. - Fit and predict temporal classifiers. - Plot and evaluate predictions. You can run these experiments using GPU with Google Colab.
1. Installing NeuralForecast
2. Loading Binary Sequence Data
Thecore.NeuralForecast class contains shared, fit, predict and
other methods that take as inputs pandas DataFrames with columns
['unique_id', 'ds', 'y'], where unique_id identifies individual time
series from the dataset, ds is the date, and y is the target binary
variable.
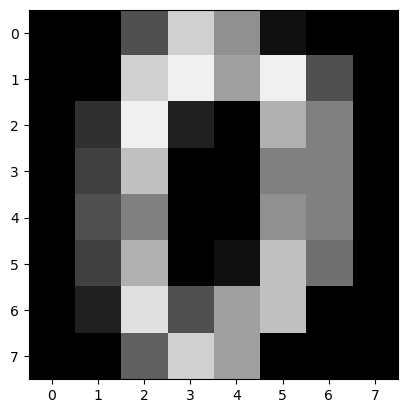
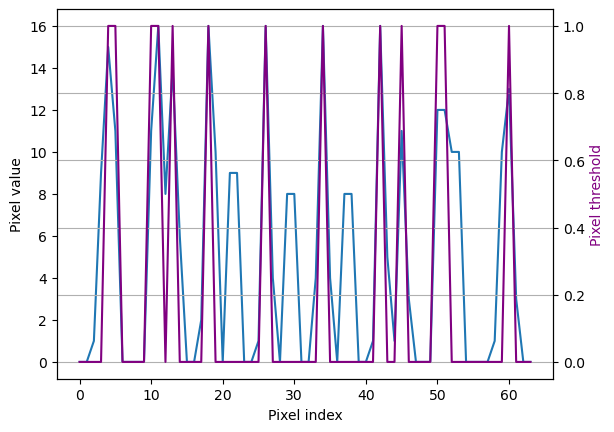
In this motivation example we convert 8x8 digits images into 64-length
sequences and define a classification problem, to identify when the
pixels surpass certain threshold. We declare a pandas dataframe in long
format, to match NeuralForecast’s inputs.
| unique_id | ds | y | pixels | |
|---|---|---|---|---|
| 0 | 0 | 1910 | 0 | 0.0 |
| 1 | 0 | 1911 | 0 | 0.0 |
| 2 | 0 | 1912 | 0 | 5.0 |
| 3 | 0 | 1913 | 1 | 13.0 |
| 4 | 0 | 1914 | 0 | 9.0 |
| … | … | … | … | … |
| 6395 | 99 | 1969 | 1 | 14.0 |
| 6396 | 99 | 1970 | 1 | 16.0 |
| 6397 | 99 | 1971 | 0 | 3.0 |
| 6398 | 99 | 1972 | 0 | 0.0 |
| 6399 | 99 | 1973 | 0 | 0.0 |
3. Fit and predict temporal classifiers
Fit the models
Using theNeuralForecast.fit method you can train a set of models to
your dataset. You can define the forecasting horizon (12 in this
example), and modify the hyperparameters of the model. For example, for
the NHITS we changed the default hidden size for both encoder and
decoders.
See the NHITS and MLP model
documentation.
Warning For the moment Recurrent-based model family is not available to operate with Bernoulli distribution output. This affects the following methodsLSTM,GRU,DilatedRNN, andTCN. This feature is work in progress.
| unique_id | ds | cutoff | MLP | MLP-median | MLP-lo-90 | MLP-lo-80 | MLP-hi-80 | MLP-hi-90 | NHITS | NHITS-median | NHITS-lo-90 | NHITS-lo-80 | NHITS-hi-80 | NHITS-hi-90 | y | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 1962 | 1961 | 0.173 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.761 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 |
| 1 | 0 | 1963 | 1961 | 0.784 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.571 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
| 2 | 0 | 1964 | 1961 | 0.042 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.009 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
| 3 | 0 | 1965 | 1961 | 0.072 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.054 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 |
| 4 | 0 | 1966 | 1961 | 0.059 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.000 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
| … | … | … | … | … | … | … | … | … | … | … | … | … | … | … | … | … |
| 1195 | 99 | 1969 | 1961 | 0.551 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.697 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
| 1196 | 99 | 1970 | 1961 | 0.662 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.465 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
| 1197 | 99 | 1971 | 1961 | 0.369 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0.382 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 |
| 1198 | 99 | 1972 | 1961 | 0.056 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0.000 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
| 1199 | 99 | 1973 | 1961 | 0.000 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.000 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
| unique_id | ds | cutoff | MLP | MLP-median | MLP-lo-90 | MLP-lo-80 | MLP-hi-80 | MLP-hi-90 | NHITS | NHITS-median | NHITS-lo-90 | NHITS-lo-80 | NHITS-hi-80 | NHITS-hi-90 | y | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0 | 1962 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 |
| 1 | 0 | 1963 | 1961 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
| 2 | 0 | 1964 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
| 3 | 0 | 1965 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 |
| 4 | 0 | 1966 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
| … | … | … | … | … | … | … | … | … | … | … | … | … | … | … | … | … |
| 1195 | 99 | 1969 | 1961 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
| 1196 | 99 | 1970 | 1961 | 1 | 1.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 1 |
| 1197 | 99 | 1971 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 1.0 | 1.0 | 0 |
| 1198 | 99 | 1972 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 1.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
| 1199 | 99 | 1973 | 1961 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 | 0.0 | 0.0 | 0.0 | 0.0 | 0.0 | 0 |
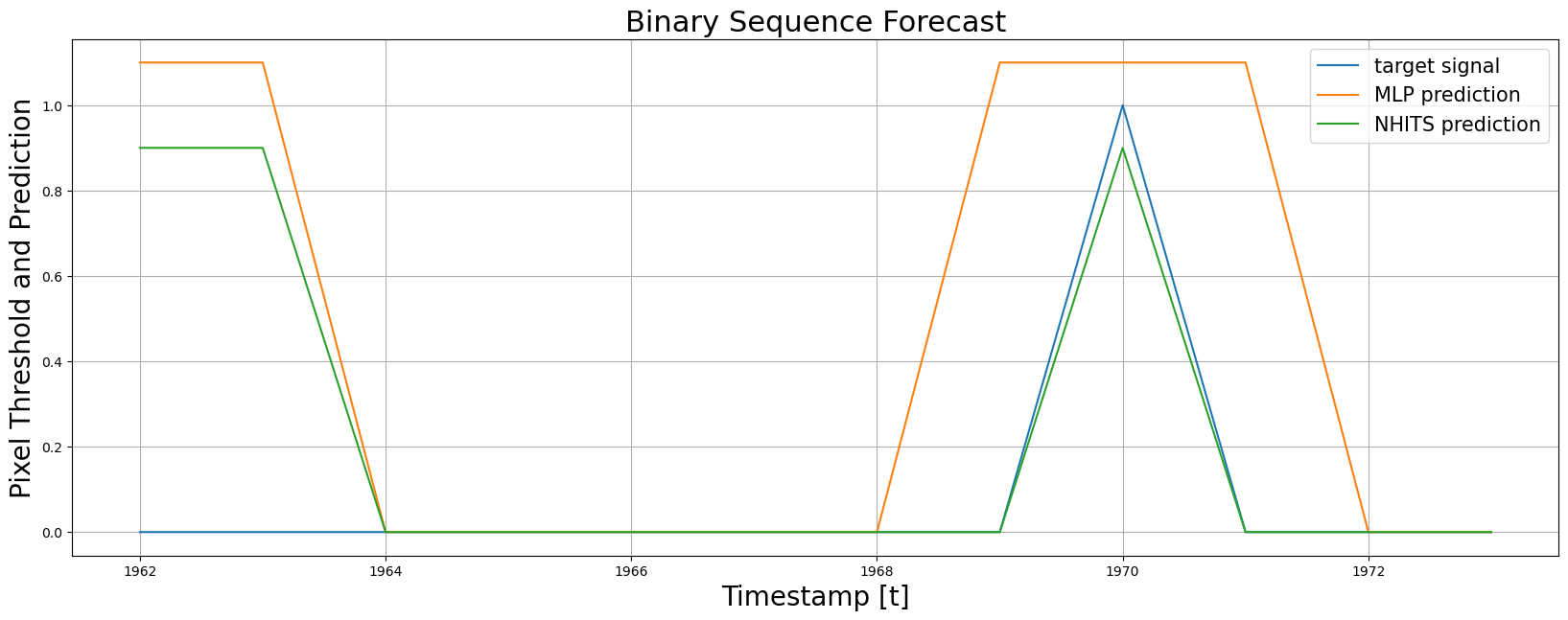
4. Plot and Evaluate Predictions
Finally, we plot the forecasts of both models against the real values. And evaluate the accuracy of theMLP and NHITS temporal classifiers.
References
- Cox D. R. (1958). “The Regression Analysis of Binary Sequences.” Journal of the Royal Statistical Society B, 20(2), 215–242.
- Cristian Challu, Kin G. Olivares, Boris N. Oreshkin, Federico Garza, Max Mergenthaler-Canseco, Artur Dubrawski (2023). NHITS: Neural Hierarchical Interpolation for Time Series Forecasting. Accepted at AAAI 2023.