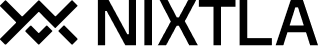Documentation Index
Fetch the complete documentation index at: https://nixtlaverse.nixtla.io/llms.txt
Use this file to discover all available pages before exploring further.
This shows an example with just 4 series of the M4 dataset. If you want
to run it yourself on all of them, you can refer to this
notebook.
import random
import lightgbm as lgb
import numpy as np
import pandas as pd
from sklearn.linear_model import LinearRegression
from utilsforecast.plotting import plot_series
from mlforecast import MLForecast
from mlforecast.lag_transforms import (
ExponentiallyWeightedMean,
RollingMean,
)
from mlforecast.lgb_cv import LightGBMCV
from mlforecast.target_transforms import Differences
from mlforecast.utils import PredictionIntervals
df = pd.read_parquet('https://datasets-nixtla.s3.amazonaws.com/m4-hourly.parquet')
ids = df['unique_id'].unique()
random.seed(0)
sample_ids = random.choices(ids, k=4)
sample_df = df[df['unique_id'].isin(sample_ids)]
sample_df
| unique_id | ds | y |
|---|
| 86796 | H196 | 1 | 11.8 |
| 86797 | H196 | 2 | 11.4 |
| 86798 | H196 | 3 | 11.1 |
| 86799 | H196 | 4 | 10.8 |
| 86800 | H196 | 5 | 10.6 |
| … | … | … | … |
| 325235 | H413 | 1004 | 99.0 |
| 325236 | H413 | 1005 | 88.0 |
| 325237 | H413 | 1006 | 47.0 |
| 325238 | H413 | 1007 | 41.0 |
| 325239 | H413 | 1008 | 34.0 |
horizon = 48
valid = sample_df.groupby('unique_id').tail(horizon)
train = sample_df.drop(valid.index)
train.shape, valid.shape
Creating the forecaster
fcst = MLForecast(
models=lgb.LGBMRegressor(random_state=0, verbosity=-1),
freq=1,
lags=[24 * (i+1) for i in range(7)],
lag_transforms={
48: [ExponentiallyWeightedMean(alpha=0.3)],
},
num_threads=1,
target_transforms=[Differences([24])],
)
MLForecast(models=[LGBMRegressor], freq=1, lag_features=['lag24', 'lag48', 'lag72', 'lag96', 'lag120', 'lag144', 'lag168', 'exponentially_weighted_mean_lag48_alpha0.3'], date_features=[], num_threads=1)
Fitting and predicting
fcst = MLForecast(
models=lgb.LGBMRegressor(random_state=0, verbosity=-1),
freq=1,
lags=[24 * (i+1) for i in range(7)],
lag_transforms={
48: [ExponentiallyWeightedMean(alpha=0.3)],
},
num_threads=1,
target_transforms=[Differences([24])],
)
train2 = train.copy()
train2['weight'] = np.random.default_rng(seed=0).random(train2.shape[0])
fcst.fit(train2, weight_col='weight', as_numpy=True).predict(5)
| unique_id | ds | LGBMRegressor |
|---|
| 0 | H196 | 961 | 16.079737 |
| 1 | H196 | 962 | 15.679737 |
| 2 | H196 | 963 | 15.279737 |
| 3 | H196 | 964 | 14.979737 |
| 4 | H196 | 965 | 14.679737 |
| 5 | H256 | 961 | 13.279737 |
| 6 | H256 | 962 | 12.679737 |
| 7 | H256 | 963 | 12.379737 |
| 8 | H256 | 964 | 12.079737 |
| 9 | H256 | 965 | 11.879737 |
| 10 | H381 | 961 | 56.939977 |
| 11 | H381 | 962 | 40.314608 |
| 12 | H381 | 963 | 33.859013 |
| 13 | H381 | 964 | 15.498139 |
| 14 | H381 | 965 | 25.722674 |
| 15 | H413 | 961 | 25.131194 |
| 16 | H413 | 962 | 19.177421 |
| 17 | H413 | 963 | 21.250829 |
| 18 | H413 | 964 | 18.743132 |
| 19 | H413 | 965 | 16.027263 |
fcst.cross_validation(train2, n_windows=2, h=5, weight_col='weight', as_numpy=True)
| unique_id | ds | cutoff | y | LGBMRegressor |
|---|
| 0 | H196 | 951 | 950 | 24.4 | 24.288850 |
| 1 | H196 | 952 | 950 | 24.3 | 24.188850 |
| 2 | H196 | 953 | 950 | 23.8 | 23.688850 |
| 3 | H196 | 954 | 950 | 22.8 | 22.688850 |
| 4 | H196 | 955 | 950 | 21.2 | 21.088850 |
| 5 | H256 | 951 | 950 | 19.5 | 19.688850 |
| 6 | H256 | 952 | 950 | 19.4 | 19.488850 |
| 7 | H256 | 953 | 950 | 18.9 | 19.088850 |
| 8 | H256 | 954 | 950 | 18.3 | 18.388850 |
| 9 | H256 | 955 | 950 | 17.0 | 17.088850 |
| 10 | H381 | 951 | 950 | 182.0 | 208.327270 |
| 11 | H381 | 952 | 950 | 222.0 | 247.768326 |
| 12 | H381 | 953 | 950 | 288.0 | 277.965997 |
| 13 | H381 | 954 | 950 | 264.0 | 321.532857 |
| 14 | H381 | 955 | 950 | 191.0 | 206.316903 |
| 15 | H413 | 951 | 950 | 77.0 | 60.972692 |
| 16 | H413 | 952 | 950 | 91.0 | 54.936494 |
| 17 | H413 | 953 | 950 | 76.0 | 73.949203 |
| 18 | H413 | 954 | 950 | 68.0 | 67.087417 |
| 19 | H413 | 955 | 950 | 68.0 | 75.896022 |
| 20 | H196 | 956 | 955 | 19.3 | 19.287891 |
| 21 | H196 | 957 | 955 | 18.2 | 18.187891 |
| 22 | H196 | 958 | 955 | 17.5 | 17.487891 |
| 23 | H196 | 959 | 955 | 16.9 | 16.887891 |
| 24 | H196 | 960 | 955 | 16.5 | 16.487891 |
| 25 | H256 | 956 | 955 | 15.5 | 15.687891 |
| 26 | H256 | 957 | 955 | 14.7 | 14.787891 |
| 27 | H256 | 958 | 955 | 14.1 | 14.287891 |
| 28 | H256 | 959 | 955 | 13.6 | 13.787891 |
| 29 | H256 | 960 | 955 | 13.2 | 13.387891 |
| 30 | H381 | 956 | 955 | 130.0 | 124.117828 |
| 31 | H381 | 957 | 955 | 113.0 | 119.180350 |
| 32 | H381 | 958 | 955 | 94.0 | 105.356552 |
| 33 | H381 | 959 | 955 | 192.0 | 127.095338 |
| 34 | H381 | 960 | 955 | 87.0 | 119.875754 |
| 35 | H413 | 956 | 955 | 59.0 | 67.993133 |
| 36 | H413 | 957 | 955 | 58.0 | 69.869815 |
| 37 | H413 | 958 | 955 | 53.0 | 34.717960 |
| 38 | H413 | 959 | 955 | 38.0 | 47.665581 |
| 39 | H413 | 960 | 955 | 46.0 | 45.940137 |
fcst.fit(train, fitted=True);
expected_future = fcst.make_future_dataframe(h=1)
expected_future
| unique_id | ds |
|---|
| 0 | H196 | 961 |
| 1 | H256 | 961 |
| 2 | H381 | 961 |
| 3 | H413 | 961 |
missing_future = fcst.get_missing_future(h=1, X_df=expected_future.head(2))
pd.testing.assert_frame_equal(
missing_future,
expected_future.tail(2).reset_index(drop=True)
)
fcst.forecast_fitted_values()
| unique_id | ds | y | LGBMRegressor |
|---|
| 0 | H196 | 193 | 12.7 | 12.671271 |
| 1 | H196 | 194 | 12.3 | 12.271271 |
| 2 | H196 | 195 | 11.9 | 11.871271 |
| 3 | H196 | 196 | 11.7 | 11.671271 |
| 4 | H196 | 197 | 11.4 | 11.471271 |
| … | … | … | … | … |
| 3067 | H413 | 956 | 59.0 | 68.280574 |
| 3068 | H413 | 957 | 58.0 | 70.427570 |
| 3069 | H413 | 958 | 53.0 | 44.767965 |
| 3070 | H413 | 959 | 38.0 | 48.691257 |
| 3071 | H413 | 960 | 46.0 | 46.652238 |
fcst.forecast_fitted_values(level=[90])
| unique_id | ds | y | LGBMRegressor | LGBMRegressor-lo-90 | LGBMRegressor-hi-90 |
|---|
| 0 | H196 | 193 | 12.7 | 12.671271 | 12.540634 | 12.801909 |
| 1 | H196 | 194 | 12.3 | 12.271271 | 12.140634 | 12.401909 |
| 2 | H196 | 195 | 11.9 | 11.871271 | 11.740634 | 12.001909 |
| 3 | H196 | 196 | 11.7 | 11.671271 | 11.540634 | 11.801909 |
| 4 | H196 | 197 | 11.4 | 11.471271 | 11.340634 | 11.601909 |
| … | … | … | … | … | … | … |
| 3067 | H413 | 956 | 59.0 | 68.280574 | 58.846640 | 77.714509 |
| 3068 | H413 | 957 | 58.0 | 70.427570 | 60.993636 | 79.861504 |
| 3069 | H413 | 958 | 53.0 | 44.767965 | 35.334031 | 54.201899 |
| 3070 | H413 | 959 | 38.0 | 48.691257 | 39.257323 | 58.125191 |
| 3071 | H413 | 960 | 46.0 | 46.652238 | 37.218304 | 56.086172 |
predictions = fcst.predict(horizon)
results = valid.merge(predictions, on=['unique_id', 'ds'])
fig = plot_series(forecasts_df=results)

Prediction intervals
With
MLForecast,
you can generate prediction intervals using Conformal Prediction. To
configure Conformal Prediction, you need to pass an instance of the
PredictionIntervals
class to the prediction_intervals argument of the fit method. The
class takes three parameters: n_windows, h and method.
n_windows represents the number of cross-validation windows used
to calibrate the intervalsh is the forecast horizonmethod can be conformal_distribution or conformal_error;
conformal_distribution (default) creates forecasts paths based on
the cross-validation errors and calculate quantiles using those
paths, on the other hand conformal_error calculates the error
quantiles to produce prediction intervals. The strategy will adjust
the intervals for each horizon step, resulting in different widths
for each step. Please note that a minimum of 2 cross-validation
windows must be used.
fcst.fit(
train,
prediction_intervals=PredictionIntervals(n_windows=3, h=48)
)
MLForecast(models=[LGBMRegressor], freq=1, lag_features=['lag24', 'lag48', 'lag72', 'lag96', 'lag120', 'lag144', 'lag168', 'exponentially_weighted_mean_lag48_alpha0.3'], date_features=[], num_threads=1)
predict method using the level argument. Levels must lie between
0 and 100.
predictions_w_intervals = fcst.predict(48, level=[50, 80, 95])
predictions_w_intervals.head()
| unique_id | ds | LGBMRegressor | LGBMRegressor-lo-95 | LGBMRegressor-lo-80 | LGBMRegressor-lo-50 | LGBMRegressor-hi-50 | LGBMRegressor-hi-80 | LGBMRegressor-hi-95 |
|---|
| 0 | H196 | 961 | 16.071271 | 15.958042 | 15.971271 | 16.005091 | 16.137452 | 16.171271 | 16.184501 |
| 1 | H196 | 962 | 15.671271 | 15.553632 | 15.553632 | 15.578632 | 15.763911 | 15.788911 | 15.788911 |
| 2 | H196 | 963 | 15.271271 | 15.153632 | 15.153632 | 15.162452 | 15.380091 | 15.388911 | 15.388911 |
| 3 | H196 | 964 | 14.971271 | 14.858042 | 14.871271 | 14.905091 | 15.037452 | 15.071271 | 15.084501 |
| 4 | H196 | 965 | 14.671271 | 14.553632 | 14.553632 | 14.562452 | 14.780091 | 14.788911 | 14.788911 |
results = valid.merge(predictions_w_intervals, on=['unique_id', 'ds'])
fig = plot_series(forecasts_df=results, level=[50, 80, 95])
fig
 If you want to reduce the computational time and produce intervals with
the same width for the whole forecast horizon, simple pass
If you want to reduce the computational time and produce intervals with
the same width for the whole forecast horizon, simple pass h=1 to the
PredictionIntervals
class. The caveat of this strategy is that in some cases, variance of
the absolute residuals maybe be small (even zero), so the intervals may
be too narrow.
fcst.fit(
train,
prediction_intervals=PredictionIntervals(n_windows=3, h=1)
);
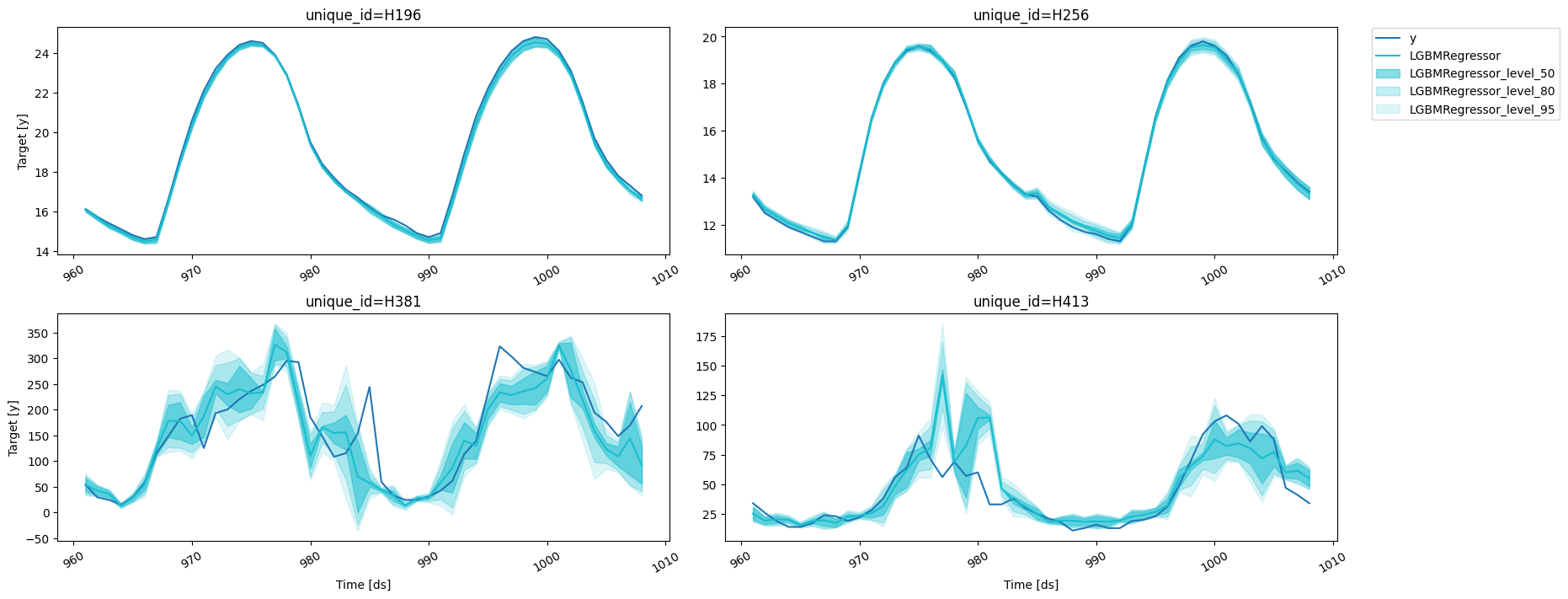
predictions_w_intervals_ws_1 = fcst.predict(48, level=[80, 90, 95])
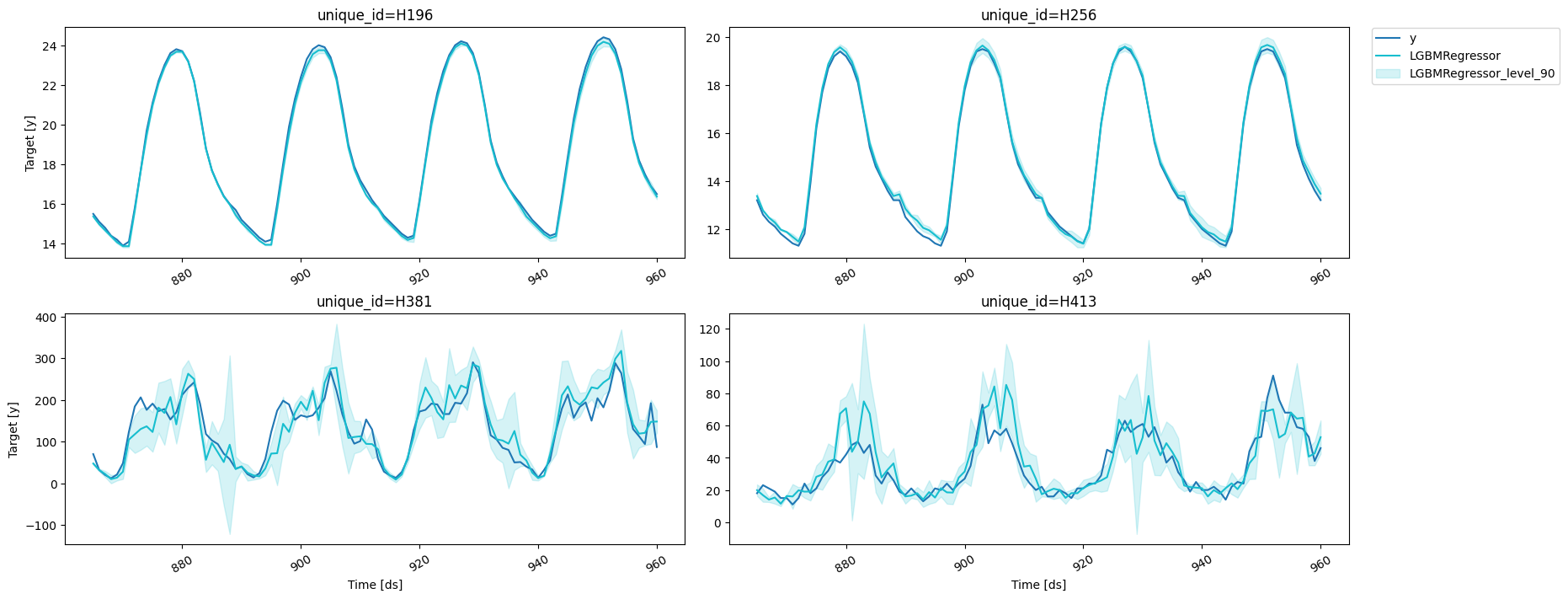
results = valid.merge(predictions_w_intervals_ws_1, on=['unique_id', 'ds'])
fig = plot_series(forecasts_df=results, level=[90])
fig
Forecast using a pretrained model
MLForecast allows you to use a pretrained model to generate forecasts
for a new dataset. Simply provide a pandas dataframe containing the new
observations as the value for the new_df argument when calling the
predict method. The dataframe should have the same structure as the
one used to fit the model, including any features and time series data.
The function will then use the pretrained model to generate forecasts
for the new observations. This allows you to easily apply a pretrained
model to a new dataset and generate forecasts without the need to
retrain the model.
ercot_df = pd.read_csv('https://datasets-nixtla.s3.amazonaws.com/ERCOT-clean.csv')
# we have to convert the ds column to integers
# since MLForecast was trained with that structure
ercot_df['ds'] = np.arange(1, len(ercot_df) + 1)
# use the `new_df` argument to pass the ercot dataset
ercot_fcsts = fcst.predict(horizon, new_df=ercot_df)
fig = plot_series(ercot_df, ercot_fcsts, max_insample_length=48 * 2)
fig

Preprocess
If you want to take a look at the data that will be used to train the
models you can call Forecast.preprocess.
prep_df = fcst.preprocess(train)
prep_df
| unique_id | ds | y | lag24 | lag48 | lag72 | lag96 | lag120 | lag144 | lag168 | exponentially_weighted_mean_lag48_alpha0.3 |
|---|
| 86988 | H196 | 193 | 0.1 | 0.0 | 0.0 | 0.0 | 0.3 | 0.1 | 0.1 | 0.3 | 0.002810 |
| 86989 | H196 | 194 | 0.1 | -0.1 | 0.1 | 0.0 | 0.3 | 0.1 | 0.1 | 0.3 | 0.031967 |
| 86990 | H196 | 195 | 0.1 | -0.1 | 0.1 | 0.0 | 0.3 | 0.1 | 0.2 | 0.1 | 0.052377 |
| 86991 | H196 | 196 | 0.1 | 0.0 | 0.0 | 0.0 | 0.3 | 0.2 | 0.1 | 0.2 | 0.036664 |
| 86992 | H196 | 197 | 0.0 | 0.0 | 0.0 | 0.1 | 0.2 | 0.2 | 0.1 | 0.2 | 0.025665 |
| … | … | … | … | … | … | … | … | … | … | … | … |
| 325187 | H413 | 956 | 0.0 | 10.0 | 1.0 | 6.0 | -53.0 | 44.0 | -21.0 | 21.0 | 7.963225 |
| 325188 | H413 | 957 | 9.0 | 10.0 | 10.0 | -7.0 | -46.0 | 27.0 | -19.0 | 24.0 | 8.574257 |
| 325189 | H413 | 958 | 16.0 | 8.0 | 5.0 | -9.0 | -36.0 | 32.0 | -13.0 | 8.0 | 7.501980 |
| 325190 | H413 | 959 | -3.0 | 17.0 | -7.0 | 2.0 | -31.0 | 22.0 | 5.0 | -2.0 | 3.151386 |
| 325191 | H413 | 960 | 15.0 | 11.0 | -6.0 | -5.0 | -17.0 | 22.0 | -18.0 | 10.0 | 0.405970 |
Forecast.fit_models, since this
only stores the series information.
X, y = prep_df.drop(columns=['unique_id', 'ds', 'y']), prep_df['y']
fcst.fit_models(X, y)
MLForecast(models=[LGBMRegressor], freq=1, lag_features=['lag24', 'lag48', 'lag72', 'lag96', 'lag120', 'lag144', 'lag168', 'exponentially_weighted_mean_lag48_alpha0.3'], date_features=[], num_threads=1)
MLForecast(models=[LGBMRegressor], freq=1, lag_features=['lag24', 'lag48', 'lag72', 'lag96', 'lag120', 'lag144', 'lag168', 'exponentially_weighted_mean_lag48_alpha0.3'], date_features=[], num_threads=1)
predictions2 = fcst.predict(horizon)
pd.testing.assert_frame_equal(predictions, predictions2)
MLForecast.cross_validation,
which takes our data and performs the process described above for
n_windows times where each window has h validation samples in it.
For example, if we have 100 samples and we want to perform 2 backtests
each of size 14, the splits will be as follows:
- Train: 1 to 72. Validation: 73 to 86.
- Train: 1 to 86. Validation: 87 to 100.
You can control the size between each cross validation window using the
step_size argument. For example, if we have 100 samples and we want to
perform 2 backtests each of size 14 and move one step ahead in each fold
(step_size=1), the splits will be as follows:
- Train: 1 to 85. Validation: 86 to 99.
- Train: 1 to 86. Validation: 87 to 100.
You can also perform cross validation without refitting your models for
each window by setting refit=False. This allows you to evaluate the
performance of your models using multiple window sizes without having to
retrain them each time.
fcst = MLForecast(
models=lgb.LGBMRegressor(random_state=0, verbosity=-1),
freq=1,
lags=[24 * (i+1) for i in range(7)],
lag_transforms={
1: [RollingMean(window_size=24)],
24: [RollingMean(window_size=24)],
48: [ExponentiallyWeightedMean(alpha=0.3)],
},
num_threads=1,
target_transforms=[Differences([24])],
)
cv_results = fcst.cross_validation(
train,
n_windows=2,
h=horizon,
step_size=horizon,
fitted=True,
)
cv_results
| unique_id | ds | cutoff | y | LGBMRegressor |
|---|
| 0 | H196 | 865 | 864 | 15.5 | 15.373393 |
| 1 | H196 | 866 | 864 | 15.1 | 14.973393 |
| 2 | H196 | 867 | 864 | 14.8 | 14.673393 |
| 3 | H196 | 868 | 864 | 14.4 | 14.373393 |
| 4 | H196 | 869 | 864 | 14.2 | 14.073393 |
| … | … | … | … | … | … |
| 379 | H413 | 956 | 912 | 59.0 | 64.284167 |
| 380 | H413 | 957 | 912 | 58.0 | 64.830429 |
| 381 | H413 | 958 | 912 | 53.0 | 40.726851 |
| 382 | H413 | 959 | 912 | 38.0 | 42.739657 |
| 383 | H413 | 960 | 912 | 46.0 | 52.802769 |
fitted=True we can access the predictions for the
training sets as well with the cross_validation_fitted_values method.
fcst.cross_validation_fitted_values()
| unique_id | ds | fold | y | LGBMRegressor |
|---|
| 0 | H196 | 193 | 0 | 12.7 | 12.673393 |
| 1 | H196 | 194 | 0 | 12.3 | 12.273393 |
| 2 | H196 | 195 | 0 | 11.9 | 11.873393 |
| 3 | H196 | 196 | 0 | 11.7 | 11.673393 |
| 4 | H196 | 197 | 0 | 11.4 | 11.473393 |
| … | … | … | … | … | … |
| 5563 | H413 | 908 | 1 | 49.0 | 50.620196 |
| 5564 | H413 | 909 | 1 | 39.0 | 35.972331 |
| 5565 | H413 | 910 | 1 | 29.0 | 29.359678 |
| 5566 | H413 | 911 | 1 | 24.0 | 25.784563 |
| 5567 | H413 | 912 | 1 | 20.0 | 23.168413 |
prediction_intervals as well as values for the width through levels.
cv_results_intervals = fcst.cross_validation(
train,
n_windows=2,
h=horizon,
step_size=horizon,
prediction_intervals=PredictionIntervals(h=horizon),
level=[80, 90]
)
cv_results_intervals
| unique_id | ds | cutoff | y | LGBMRegressor | LGBMRegressor-lo-90 | LGBMRegressor-lo-80 | LGBMRegressor-hi-80 | LGBMRegressor-hi-90 |
|---|
| 0 | H196 | 865 | 864 | 15.5 | 15.373393 | 15.311379 | 15.316528 | 15.430258 | 15.435407 |
| 1 | H196 | 866 | 864 | 15.1 | 14.973393 | 14.940556 | 14.940556 | 15.006230 | 15.006230 |
| 2 | H196 | 867 | 864 | 14.8 | 14.673393 | 14.606230 | 14.606230 | 14.740556 | 14.740556 |
| 3 | H196 | 868 | 864 | 14.4 | 14.373393 | 14.306230 | 14.306230 | 14.440556 | 14.440556 |
| 4 | H196 | 869 | 864 | 14.2 | 14.073393 | 14.006230 | 14.006230 | 14.140556 | 14.140556 |
| … | … | … | … | … | … | … | … | … | … |
| 379 | H413 | 956 | 912 | 59.0 | 64.284167 | 29.890099 | 34.371545 | 94.196788 | 98.678234 |
| 380 | H413 | 957 | 912 | 58.0 | 64.830429 | 56.874572 | 57.827689 | 71.833169 | 72.786285 |
| 381 | H413 | 958 | 912 | 53.0 | 40.726851 | 35.296195 | 35.846206 | 45.607495 | 46.157506 |
| 382 | H413 | 959 | 912 | 38.0 | 42.739657 | 35.292153 | 35.807640 | 49.671674 | 50.187161 |
| 383 | H413 | 960 | 912 | 46.0 | 52.802769 | 42.465597 | 43.895670 | 61.709869 | 63.139941 |
refit argument allows us to control if we want to retrain the
models in every window. It can either be:
- A boolean: True will retrain on every window and False only on the
first one.
- A positive integer: The models will be trained on the first window
and then every
refit windows.
fcst = MLForecast(
models=LinearRegression(),
freq=1,
lags=[1, 24],
)
for refit, expected_models in zip([True, False, 2], [4, 1, 2]):
fcst.cross_validation(
train,
n_windows=4,
h=horizon,
refit=refit,
)
assert len(fcst.cv_models_) == expected_models
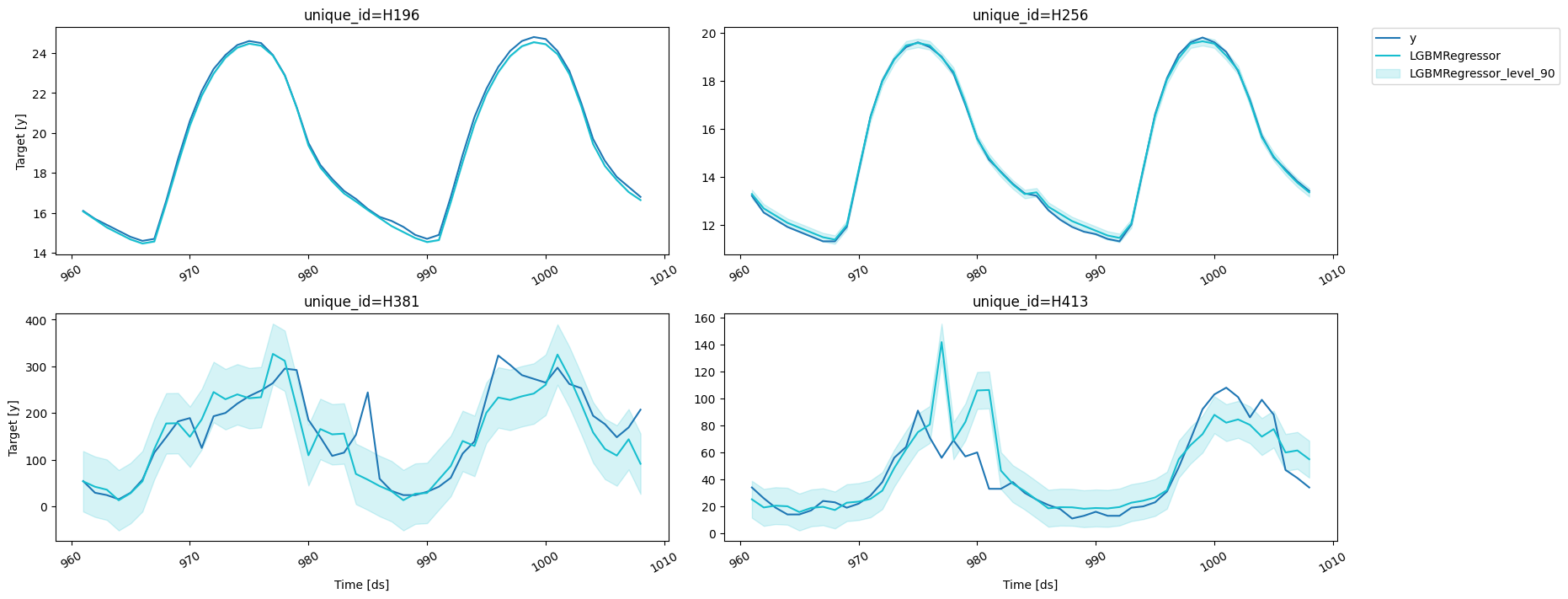
fig = plot_series(forecasts_df=cv_results.drop(columns='cutoff'))
fig
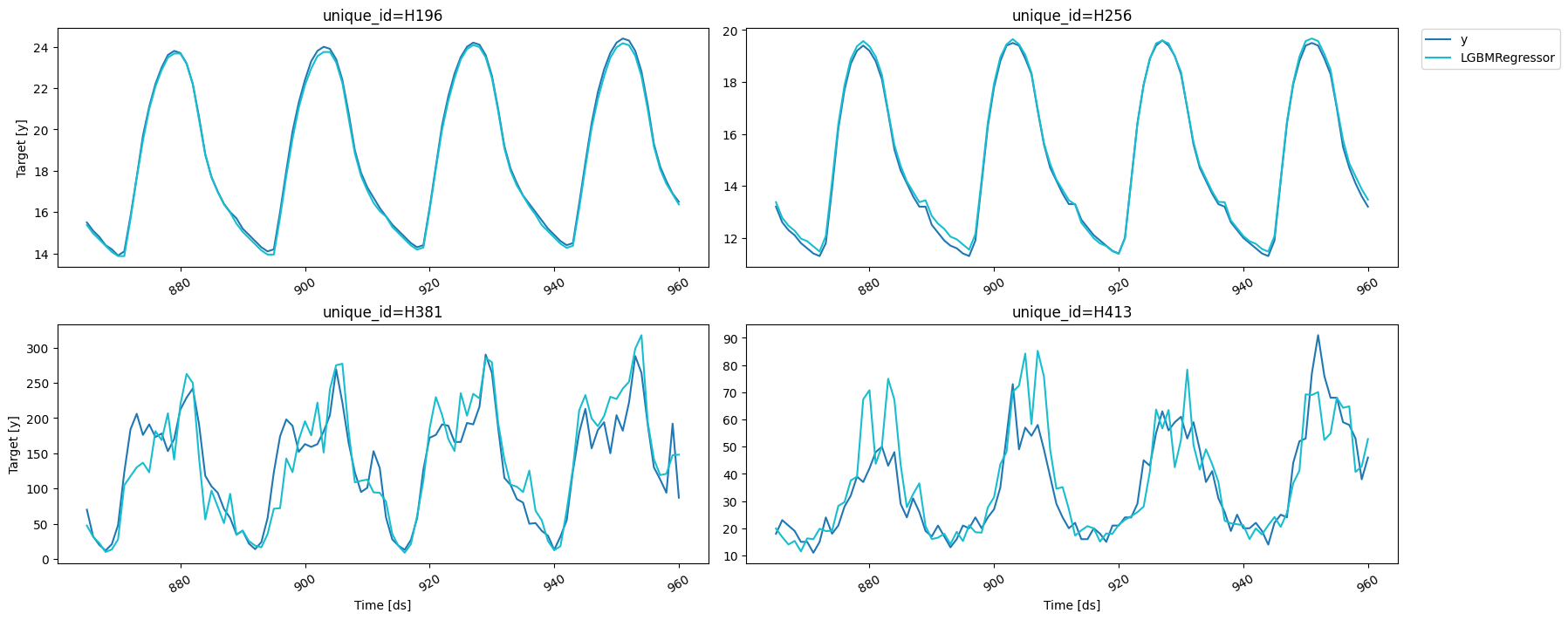
fig = plot_series(forecasts_df=cv_results_intervals.drop(columns='cutoff'), level=[90])
fig
Using LightGBMCV to tune your forecasts
Once you’ve found a set of features and parameters that work for your
problem you can build a forecast object from it using
MLForecast.from_cv,
which takes the trained
LightGBMCV
object and builds an
MLForecast
object that will use the same features and parameters. Then you can call
fit and predict as you normally would.
cv = LightGBMCV(
freq=1,
lags=[24 * (i+1) for i in range(7)],
lag_transforms={
48: [ExponentiallyWeightedMean(alpha=0.3)],
},
num_threads=1,
target_transforms=[Differences([24])]
)
hist = cv.fit(
train,
n_windows=2,
h=horizon,
params={'verbosity': -1},
)
[10] mape: 0.118569
[20] mape: 0.111506
[30] mape: 0.107314
[40] mape: 0.106089
[50] mape: 0.106630
Early stopping at round 50
Using best iteration: 40
fcst = MLForecast.from_cv(cv)
assert cv.best_iteration_ == fcst.models['LGBMRegressor'].n_estimators