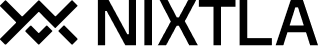Documentation Index
Fetch the complete documentation index at: https://nixtlaverse.nixtla.io/llms.txt
Use this file to discover all available pages before exploring further.
Detailed description of all the functionalities that MLForecast
provides.
Data setup
For this example we’ll use a subset of the M4 hourly dataset. You can
find the a notebook with the full dataset
here.
import random
import tempfile
from pathlib import Path
import pandas as pd
from datasetsforecast.m4 import M4
from utilsforecast.plotting import plot_series
await M4.async_download('data', group='Hourly')
df, *_ = M4.load('data', 'Hourly')
uids = df['unique_id'].unique()
random.seed(0)
sample_uids = random.choices(uids, k=4)
df = df[df['unique_id'].isin(sample_uids)].reset_index(drop=True)
df['ds'] = df['ds'].astype('int64')
df
| unique_id | ds | y |
|---|
| 0 | H196 | 1 | 11.8 |
| 1 | H196 | 2 | 11.4 |
| 2 | H196 | 3 | 11.1 |
| 3 | H196 | 4 | 10.8 |
| 4 | H196 | 5 | 10.6 |
| … | … | … | … |
| 4027 | H413 | 1004 | 99.0 |
| 4028 | H413 | 1005 | 88.0 |
| 4029 | H413 | 1006 | 47.0 |
| 4030 | H413 | 1007 | 41.0 |
| 4031 | H413 | 1008 | 34.0 |
EDA
We’ll take a look at our series to get ideas for transformations and
features.
fig = plot_series(df, max_insample_length=24 * 14)
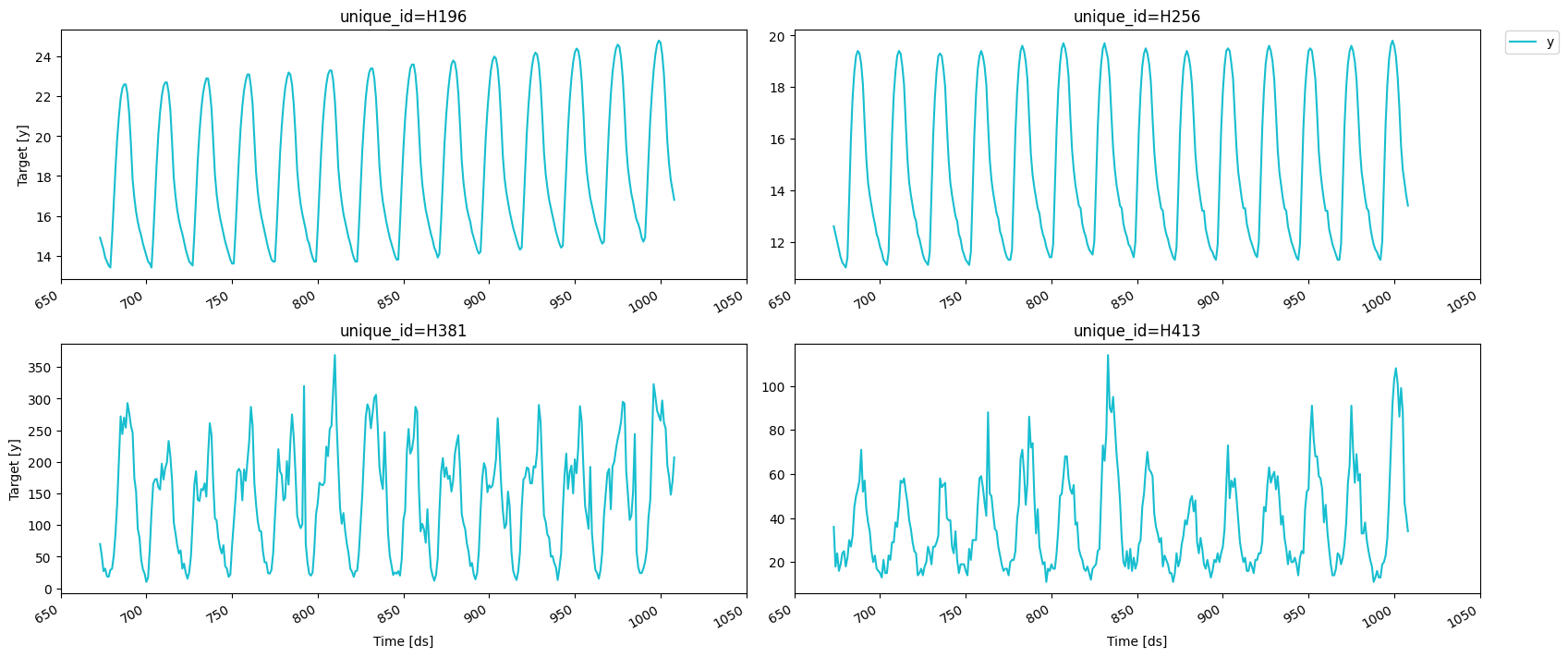 We can use the
We can use the MLForecast.preprocess method to explore different
transformations. It looks like these series have a strong seasonality on
the hour of the day, so we can subtract the value from the same hour in
the previous day to remove it. This can be done with the
mlforecast.target_transforms.Differences transformer, which we pass
through target_transforms.
from mlforecast import MLForecast
from mlforecast.target_transforms import Differences
fcst = MLForecast(
models=[], # we're not interested in modeling yet
freq=1, # our series have integer timestamps, so we'll just add 1 in every timestep
target_transforms=[Differences([24])],
)
prep = fcst.preprocess(df)
prep
| unique_id | ds | y |
|---|
| 24 | H196 | 25 | 0.3 |
| 25 | H196 | 26 | 0.3 |
| 26 | H196 | 27 | 0.1 |
| 27 | H196 | 28 | 0.2 |
| 28 | H196 | 29 | 0.2 |
| … | … | … | … |
| 4027 | H413 | 1004 | 39.0 |
| 4028 | H413 | 1005 | 55.0 |
| 4029 | H413 | 1006 | 14.0 |
| 4030 | H413 | 1007 | 3.0 |
| 4031 | H413 | 1008 | 4.0 |
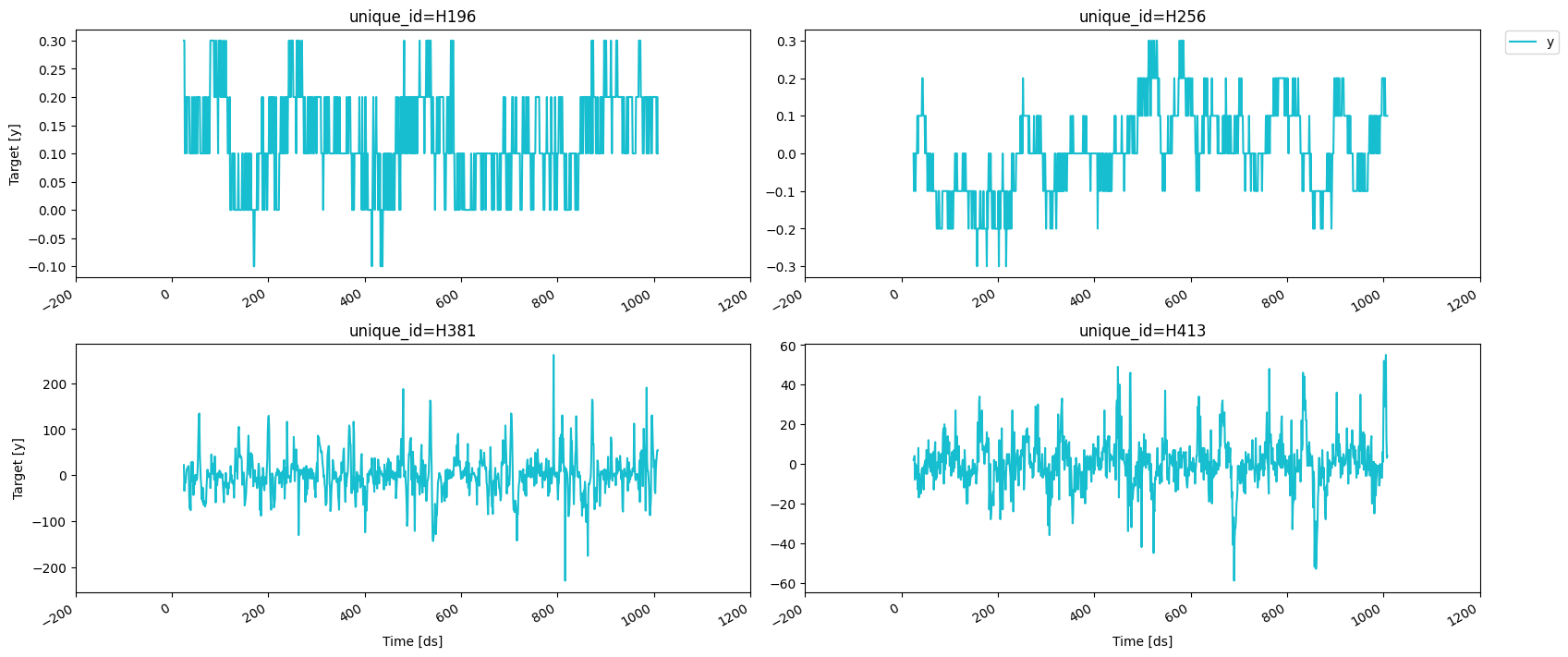
Adding features
Lags
Looks like the seasonality is gone, we can now try adding some lag
features.
fcst = MLForecast(
models=[],
freq=1,
lags=[1, 24],
target_transforms=[Differences([24])],
)
prep = fcst.preprocess(df)
prep
| unique_id | ds | y | lag1 | lag24 |
|---|
| 48 | H196 | 49 | 0.1 | 0.1 | 0.3 |
| 49 | H196 | 50 | 0.1 | 0.1 | 0.3 |
| 50 | H196 | 51 | 0.2 | 0.1 | 0.1 |
| 51 | H196 | 52 | 0.1 | 0.2 | 0.2 |
| 52 | H196 | 53 | 0.1 | 0.1 | 0.2 |
| … | … | … | … | … | … |
| 4027 | H413 | 1004 | 39.0 | 29.0 | 1.0 |
| 4028 | H413 | 1005 | 55.0 | 39.0 | -25.0 |
| 4029 | H413 | 1006 | 14.0 | 55.0 | -20.0 |
| 4030 | H413 | 1007 | 3.0 | 14.0 | 0.0 |
| 4031 | H413 | 1008 | 4.0 | 3.0 | -16.0 |
prep.drop(columns=['unique_id', 'ds']).corr()['y']
y 1.000000
lag1 0.622531
lag24 -0.234268
Name: y, dtype: float64
mlforecast.lag_transforms module or numba
jitted functions (so that computing the features doesn’t become a
bottleneck and we can bypass the GIL when using multithreading), we have
some implemented in the window-ops
package but you can also
implement your own.
from mlforecast.lag_transforms import ExpandingMean, RollingMean
from numba import njit
from window_ops.rolling import rolling_mean
@njit
def rolling_mean_48(x):
return rolling_mean(x, window_size=48)
fcst = MLForecast(
models=[],
freq=1,
target_transforms=[Differences([24])],
lag_transforms={
1: [ExpandingMean()],
24: [RollingMean(window_size=48), rolling_mean_48],
},
)
prep = fcst.preprocess(df)
prep
| unique_id | ds | y | expanding_mean_lag1 | rolling_mean_lag24_window_size48 | rolling_mean_48_lag24 |
|---|
| 95 | H196 | 96 | 0.1 | 0.174648 | 0.150000 | 0.150000 |
| 96 | H196 | 97 | 0.3 | 0.173611 | 0.145833 | 0.145833 |
| 97 | H196 | 98 | 0.3 | 0.175342 | 0.141667 | 0.141667 |
| 98 | H196 | 99 | 0.3 | 0.177027 | 0.141667 | 0.141667 |
| 99 | H196 | 100 | 0.3 | 0.178667 | 0.141667 | 0.141667 |
| … | … | … | … | … | … | … |
| 4027 | H413 | 1004 | 39.0 | 0.242084 | 3.437500 | 3.437500 |
| 4028 | H413 | 1005 | 55.0 | 0.281633 | 2.708333 | 2.708333 |
| 4029 | H413 | 1006 | 14.0 | 0.337411 | 2.125000 | 2.125000 |
| 4030 | H413 | 1007 | 3.0 | 0.351324 | 1.770833 | 1.770833 |
| 4031 | H413 | 1008 | 4.0 | 0.354018 | 1.208333 | 1.208333 |
Date features
If your time column is made of timestamps then it might make sense to
extract features like week, dayofweek, quarter, etc. You can do that by
passing a list of strings with pandas time/date
components.
You can also pass functions that will take the time column as input, as
we’ll show here.
def hour_index(times):
return times % 24
fcst = MLForecast(
models=[],
freq=1,
target_transforms=[Differences([24])],
date_features=[hour_index],
)
fcst.preprocess(df)
| unique_id | ds | y | hour_index |
|---|
| 24 | H196 | 25 | 0.3 | 1 |
| 25 | H196 | 26 | 0.3 | 2 |
| 26 | H196 | 27 | 0.1 | 3 |
| 27 | H196 | 28 | 0.2 | 4 |
| 28 | H196 | 29 | 0.2 | 5 |
| … | … | … | … | … |
| 4027 | H413 | 1004 | 39.0 | 20 |
| 4028 | H413 | 1005 | 55.0 | 21 |
| 4029 | H413 | 1006 | 14.0 | 22 |
| 4030 | H413 | 1007 | 3.0 | 23 |
| 4031 | H413 | 1008 | 4.0 | 0 |
target_transforms argument, which takes a list of transformations. You
can find the implemented ones in mlforecast.target_transforms or you
can implement your own as described in the target transformations
guide.
from mlforecast.target_transforms import LocalStandardScaler
fcst = MLForecast(
models=[],
freq=1,
lags=[1],
target_transforms=[LocalStandardScaler()]
)
fcst.preprocess(df)
| unique_id | ds | y | lag1 |
|---|
| 1 | H196 | 2 | -1.493026 | -1.383286 |
| 2 | H196 | 3 | -1.575331 | -1.493026 |
| 3 | H196 | 4 | -1.657635 | -1.575331 |
| 4 | H196 | 5 | -1.712505 | -1.657635 |
| 5 | H196 | 6 | -1.794810 | -1.712505 |
| … | … | … | … | … |
| 4027 | H413 | 1004 | 3.062766 | 2.425012 |
| 4028 | H413 | 1005 | 2.523128 | 3.062766 |
| 4029 | H413 | 1006 | 0.511751 | 2.523128 |
| 4030 | H413 | 1007 | 0.217403 | 0.511751 |
| 4031 | H413 | 1008 | -0.126003 | 0.217403 |
from sklearn.base import BaseEstimator
class Naive(BaseEstimator):
def fit(self, X, y):
return self
def predict(self, X):
return X['lag1']
fcst = MLForecast(
models=[Naive()],
freq=1,
lags=[1],
target_transforms=[LocalStandardScaler()]
)
fcst.fit(df)
preds = fcst.predict(1)
preds
| unique_id | ds | Naive |
|---|
| 0 | H196 | 1009 | 16.8 |
| 1 | H256 | 1009 | 13.4 |
| 2 | H381 | 1009 | 207.0 |
| 3 | H413 | 1009 | 34.0 |
last_vals = df.groupby('unique_id').tail(1)
last_vals
| unique_id | ds | y |
|---|
| 1007 | H196 | 1008 | 16.8 |
| 2015 | H256 | 1008 | 13.4 |
| 3023 | H381 | 1008 | 207.0 |
| 4031 | H413 | 1008 | 34.0 |
np.testing.assert_allclose(preds['Naive'], last_vals['y'])
Training
Once you’ve decided the features, transformations and models that you
want to use you can use the MLForecast.fit method instead, which will
do the preprocessing and then train the models. The models can be
specified as a list (which will name them by using their class name and
an index if there are repeated classes) or as a dictionary where the
keys are the names you want to give to the models, i.e. the name of the
column that will hold their predictions, and the values are the models
themselves.
lgb_params = {
'verbosity': -1,
'num_leaves': 512,
}
fcst = MLForecast(
models={
'avg': lgb.LGBMRegressor(**lgb_params),
'q75': lgb.LGBMRegressor(**lgb_params, objective='quantile', alpha=0.75),
'q25': lgb.LGBMRegressor(**lgb_params, objective='quantile', alpha=0.25),
},
freq=1,
target_transforms=[Differences([24])],
lags=[1, 24],
lag_transforms={
1: [ExpandingMean()],
24: [RollingMean(window_size=48)],
},
date_features=[hour_index],
)
fcst.fit(df)
MLForecast(models=[avg, q75, q25], freq=1, lag_features=['lag1', 'lag24', 'expanding_mean_lag1', 'rolling_mean_lag24_window_size48'], date_features=[<function hour_index>], num_threads=1)
Forecasting
preds = fcst.predict(48)
preds
| unique_id | ds | avg | q75 | q25 |
|---|
| 0 | H196 | 1009 | 16.295257 | 16.357148 | 16.315731 |
| 1 | H196 | 1010 | 15.910282 | 16.007322 | 15.862261 |
| 2 | H196 | 1011 | 15.728367 | 15.780183 | 15.658180 |
| 3 | H196 | 1012 | 15.468414 | 15.513598 | 15.399717 |
| 4 | H196 | 1013 | 15.081279 | 15.133848 | 15.007694 |
| … | … | … | … | … | … |
| 187 | H413 | 1052 | 100.450617 | 124.211150 | 47.025017 |
| 188 | H413 | 1053 | 88.426800 | 108.303409 | 44.715380 |
| 189 | H413 | 1054 | 59.675737 | 81.859964 | 19.239462 |
| 190 | H413 | 1055 | 57.580356 | 72.703301 | 21.486674 |
| 191 | H413 | 1056 | 42.669879 | 46.018271 | 24.392357 |
fig = plot_series(df, preds, max_insample_length=24 * 7)
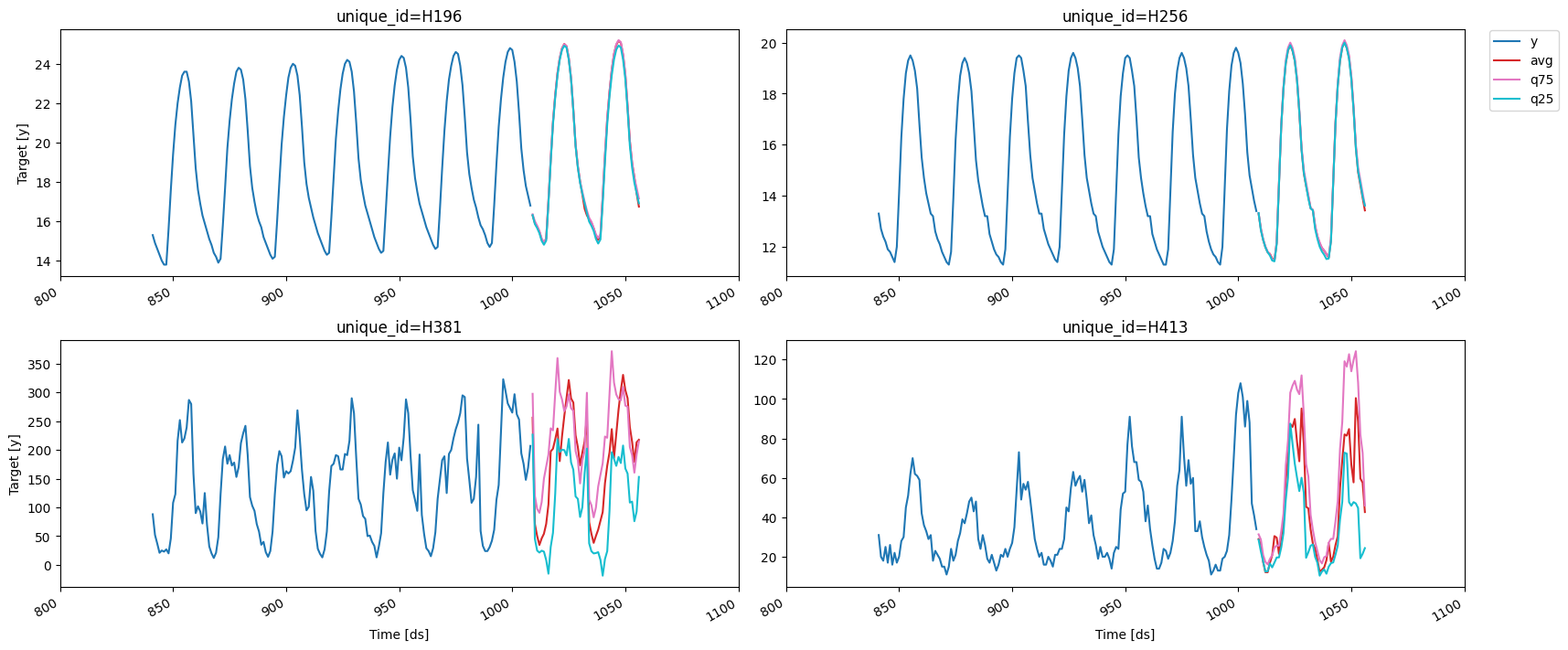
Saving and loading
The MLForecast class has the MLForecast.save and MLForecast.load to
store and then load the forecast object.
with tempfile.TemporaryDirectory() as tmpdir:
save_dir = Path(tmpdir) / 'mlforecast'
fcst.save(save_dir)
fcst2 = MLForecast.load(save_dir)
preds2 = fcst2.predict(48)
pd.testing.assert_frame_equal(preds, preds2)
Updating series’ values
After you’ve trained a forecast object you can save and load it with the
previous methods. If by the time you want to use it you already know the
following values of the target you can use the MLForecast.update
method to incorporate these, which will allow you to use these new
values when computing predictions.
- If no new values are provided for a series that’s currently stored,
only the previous ones are kept.
- If new series are included they are added to the existing ones.
fcst = MLForecast(
models=[Naive()],
freq=1,
lags=[1, 2, 3],
)
fcst.fit(df)
fcst.predict(1)
| unique_id | ds | Naive |
|---|
| 0 | H196 | 1009 | 16.8 |
| 1 | H256 | 1009 | 13.4 |
| 2 | H381 | 1009 | 207.0 |
| 3 | H413 | 1009 | 34.0 |
new_values = pd.DataFrame({
'unique_id': ['H196', 'H256'],
'ds': [1009, 1009],
'y': [17.0, 14.0],
})
fcst.update(new_values)
preds = fcst.predict(1)
preds
| unique_id | ds | Naive |
|---|
| 0 | H196 | 1010 | 17.0 |
| 1 | H256 | 1010 | 14.0 |
| 2 | H381 | 1009 | 207.0 |
| 3 | H413 | 1009 | 34.0 |
Cross validation
In order to get an estimate of how well our model will be when
predicting future data we can perform cross validation, which consists
of training a few models independently on different subsets of the data,
using them to predict a validation set and measuring their performance.
Since our data depends on time, we make our splits by removing the last
portions of the series and using them as validation sets. This process
is implemented in MLForecast.cross_validation.
fcst = MLForecast(
models=lgb.LGBMRegressor(**lgb_params),
freq=1,
target_transforms=[Differences([24])],
lags=[1, 24],
lag_transforms={
1: [ExpandingMean()],
24: [RollingMean(window_size=48)],
},
date_features=[hour_index],
)
cv_result = fcst.cross_validation(
df,
n_windows=4, # number of models to train/splits to perform
h=48, # length of the validation set in each window
)
cv_result
| unique_id | ds | cutoff | y | LGBMRegressor |
|---|
| 0 | H196 | 817 | 816 | 15.3 | 15.383165 |
| 1 | H196 | 818 | 816 | 14.9 | 14.923219 |
| 2 | H196 | 819 | 816 | 14.6 | 14.667834 |
| 3 | H196 | 820 | 816 | 14.2 | 14.275964 |
| 4 | H196 | 821 | 816 | 13.9 | 13.973491 |
| … | … | … | … | … | … |
| 763 | H413 | 1004 | 960 | 99.0 | 65.644823 |
| 764 | H413 | 1005 | 960 | 88.0 | 71.717097 |
| 765 | H413 | 1006 | 960 | 47.0 | 76.704377 |
| 766 | H413 | 1007 | 960 | 41.0 | 53.446638 |
| 767 | H413 | 1008 | 960 | 34.0 | 54.902634 |
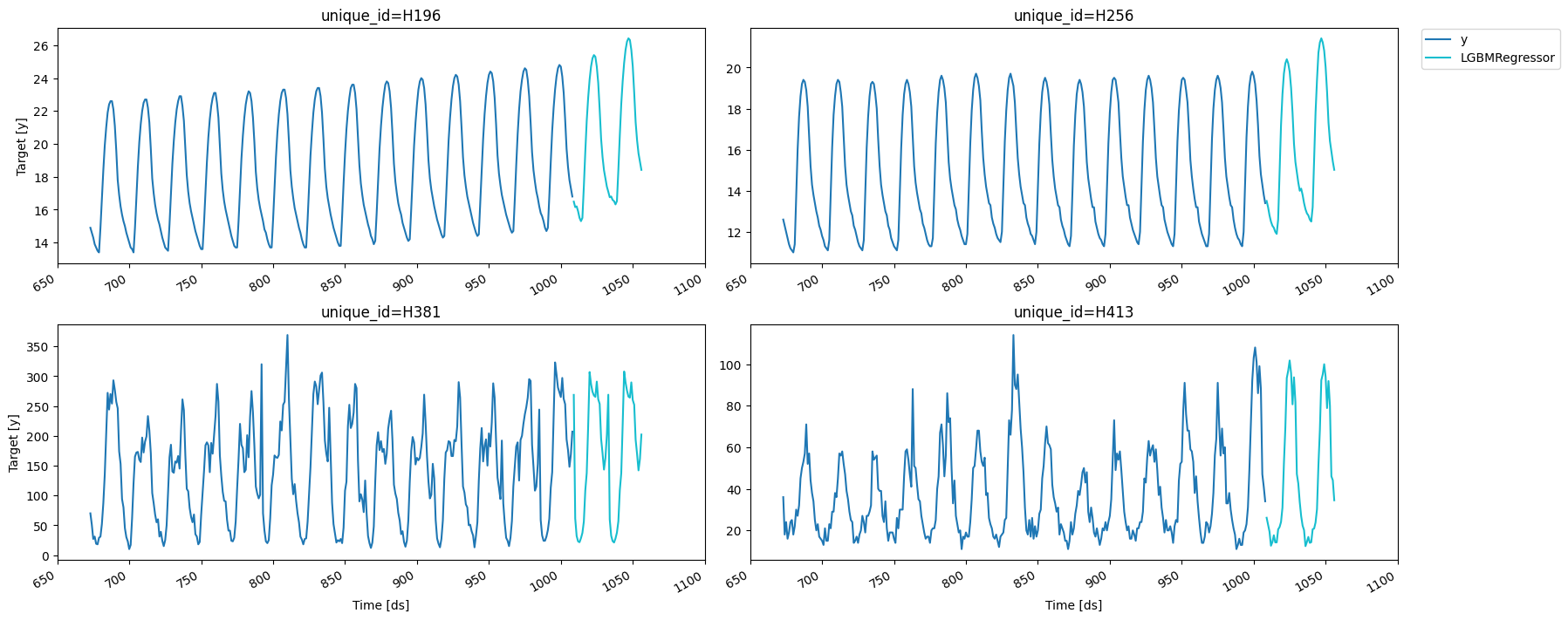
fig = plot_series(forecasts_df=cv_result.drop(columns='cutoff'))
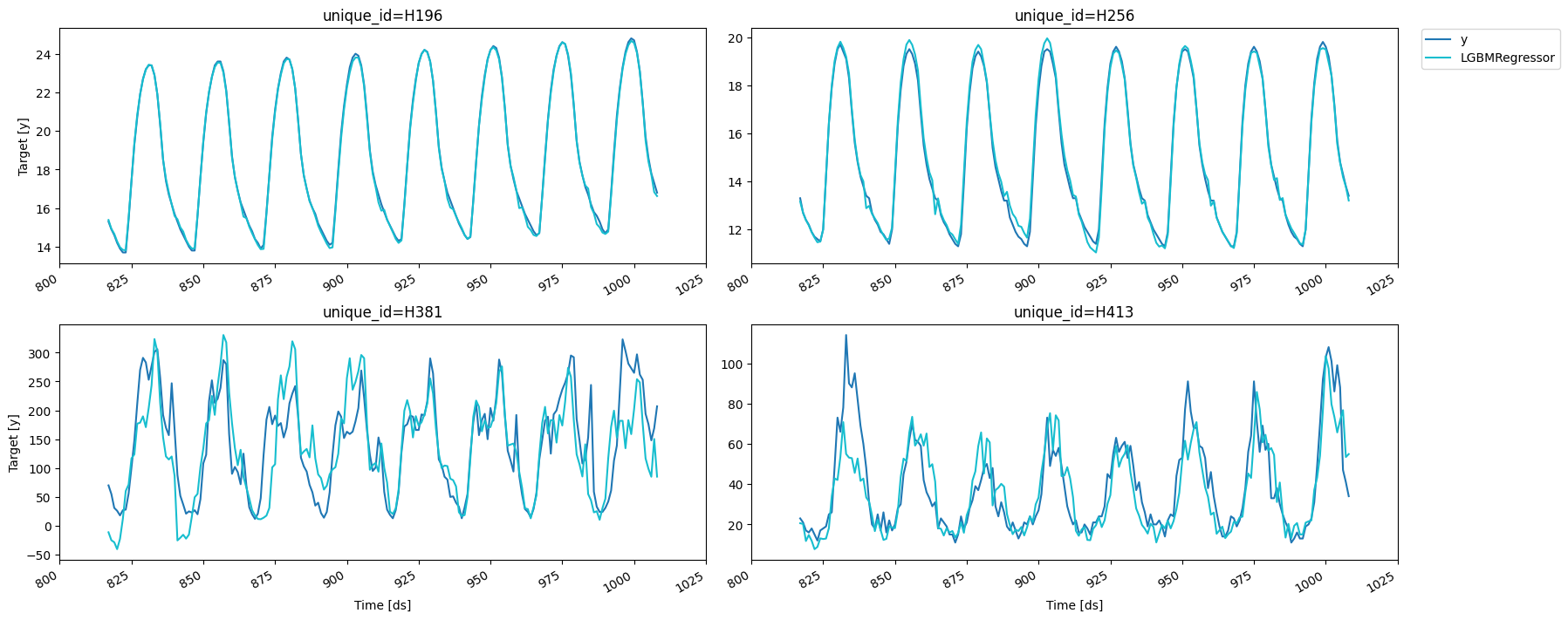 We can compute the RMSE on each split.
We can compute the RMSE on each split.
from utilsforecast.losses import rmse
def evaluate_cv(df):
return rmse(df, models=['LGBMRegressor'], id_col='cutoff').set_index('cutoff')
split_rmse = evaluate_cv(cv_result)
split_rmse
| LGBMRegressor |
|---|
| cutoff | |
| 816 | 29.418172 |
| 864 | 34.257598 |
| 912 | 13.145763 |
| 960 | 35.066261 |
LGBMRegressor 27.971949
dtype: float64
from mlforecast.lag_transforms import ExponentiallyWeightedMean
fcst = MLForecast(
models=lgb.LGBMRegressor(**lgb_params),
freq=1,
lags=[1, 24],
lag_transforms={
1: [ExponentiallyWeightedMean(alpha=0.5)],
24: [RollingMean(window_size=48)],
},
date_features=[hour_index],
)
cv_result2 = fcst.cross_validation(
df,
n_windows=4,
h=48,
)
evaluate_cv(cv_result2).mean()
LGBMRegressor 25.874439
dtype: float64
LightGBMCV
In the same spirit of estimating our model’s performance, LightGBMCV
allows us to train a few
LightGBM models on different
partitions of the data. The main differences with
MLForecast.cross_validation are:
- It can only train LightGBM models.
- It trains all models simultaneously and gives us per-iteration
averages of the errors across the complete forecasting window, which
allows us to find the best iteration.
from mlforecast.lgb_cv import LightGBMCV
cv = LightGBMCV(
freq=1,
target_transforms=[Differences([24])],
lags=[1, 24],
lag_transforms={
1: [ExpandingMean()],
24: [RollingMean(window_size=48)],
},
date_features=[hour_index],
num_threads=2,
)
cv_hist = cv.fit(
df,
n_windows=4,
h=48,
params=lgb_params,
eval_every=5,
early_stopping_evals=5,
compute_cv_preds=True,
)
[5] mape: 0.158639
[10] mape: 0.163739
[15] mape: 0.161535
[20] mape: 0.169491
[25] mape: 0.163690
[30] mape: 0.164198
Early stopping at round 30
Using best iteration: 5
eval_every argument) and performs early stopping (which can be
configured with early_stopping_evals and early_stopping_pct). If you
set compute_cv_preds=True the out-of-fold predictions are computed
using the best iteration found and are saved in the cv_preds_
attribute.
| unique_id | ds | y | Booster | window |
|---|
| 0 | H196 | 817 | 15.3 | 15.473182 | 0 |
| 1 | H196 | 818 | 14.9 | 15.038571 | 0 |
| 2 | H196 | 819 | 14.6 | 14.849409 | 0 |
| 3 | H196 | 820 | 14.2 | 14.448379 | 0 |
| 4 | H196 | 821 | 13.9 | 14.148379 | 0 |
| … | … | … | … | … | … |
| 187 | H413 | 1004 | 99.0 | 61.425396 | 3 |
| 188 | H413 | 1005 | 88.0 | 62.886890 | 3 |
| 189 | H413 | 1006 | 47.0 | 57.886890 | 3 |
| 190 | H413 | 1007 | 41.0 | 38.849009 | 3 |
| 191 | H413 | 1008 | 34.0 | 44.720562 | 3 |
fig = plot_series(forecasts_df=cv.cv_preds_.drop(columns='window'))
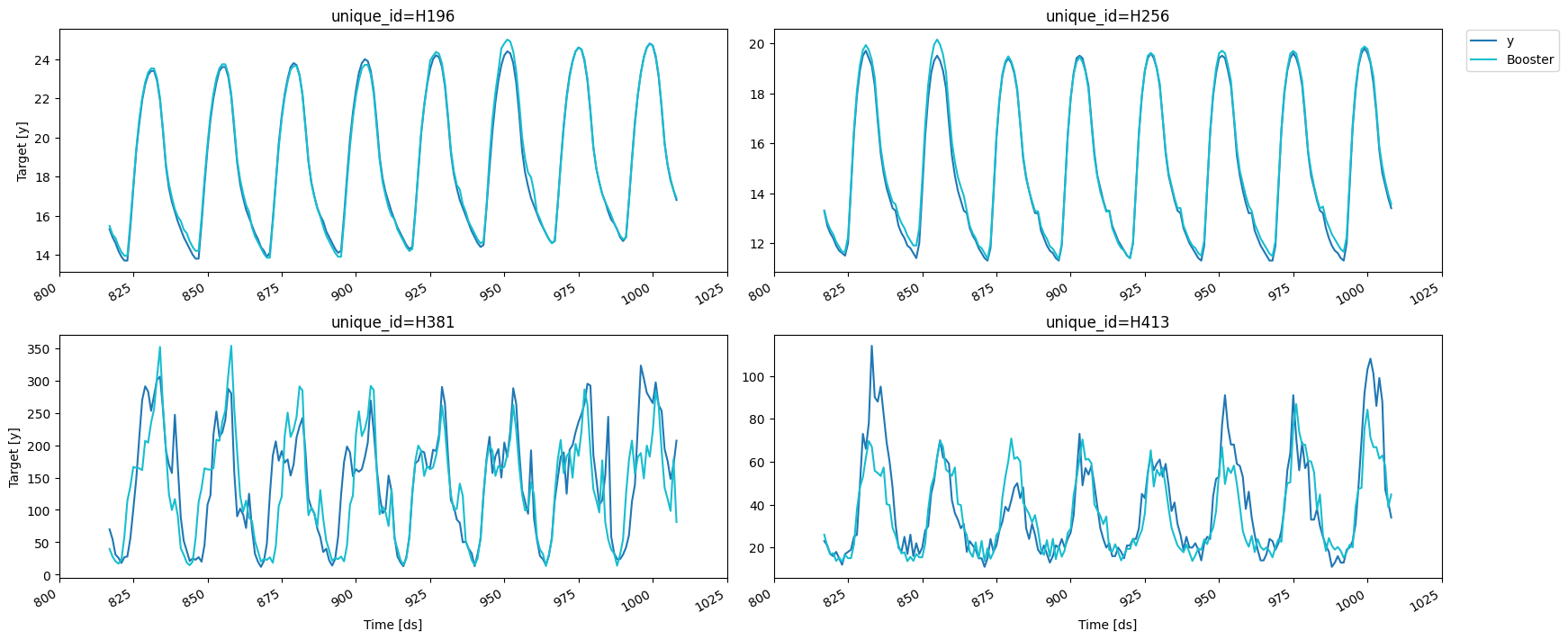 You can use this class to quickly try different configurations of
features and hyperparameters. Once you’ve found a combination that works
you can train a model with those features and hyperparameters on all the
data by creating an
You can use this class to quickly try different configurations of
features and hyperparameters. Once you’ve found a combination that works
you can train a model with those features and hyperparameters on all the
data by creating an MLForecast object from the LightGBMCV one as
follows:
final_fcst = MLForecast.from_cv(cv)
final_fcst.fit(df)
preds = final_fcst.predict(48)
fig = plot_series(df, preds, max_insample_length=24 * 14)