Learn how to select the best forecasting models when you care about more than one metric — without manually ranking and comparing every combination.
What you’ll learn
- Why single-metric model selection can be misleading
- What Pareto dominance means and when a model is “dominated”
- How to use
evaluate()correctly as input toParetoFrontier - How to visualize the 2D Pareto frontier across any two metrics
- How to handle cross-validation output for multi-objective selection
The problem: picking one winner across multiple metrics
After training several forecasting models, a common question is: which one should I deploy? If you optimize for a single metric — say MAE — the answer is straightforward: pick the lowest MAE. But real-world requirements rarely reduce to a single number. You might care about:- Accuracy (MAE, RMSE): how close are point forecasts to actuals?
- Relative error (MAPE, sMAPE): how large is the error relative to the scale of the series?
- Bias (bias, CFE): does the model systematically over- or under-forecast?
Install libraries
Generate synthetic time series
We usegenerate_series to create a panel of daily time series with
weekly seasonality. Each series contains between 100 and 150
observations, giving models enough history for a meaningful fit.
| unique_id | ds | y | |
|---|---|---|---|
| 0 | 0 | 2000-01-01 | 0.049987 |
| 1 | 0 | 2000-01-02 | 1.229624 |
| 2 | 0 | 2000-01-03 | 2.166854 |
| 3 | 0 | 2000-01-04 | 3.071433 |
| 4 | 0 | 2000-01-05 | 4.325444 |
Fit models and generate forecasts
We compare three models that cover a range of complexity:- SeasonalNaive — repeats the last observed season. Fast, transparent, surprisingly hard to beat.
- AutoARIMA — fits a SARIMA model selected automatically by AIC. More flexible but slower.
- MSTL — decomposes the series into trend and seasonal components using STL, then forecasts each part separately. Good at capturing multiple seasonal patterns.
| unique_id | ds | SeasonalNaive | AutoARIMA | MSTL | |
|---|---|---|---|---|---|
| 0 | 0 | 2000-05-04 | 5.453414 | 5.296218 | 5.283464 |
| 1 | 0 | 2000-05-05 | 6.136066 | 6.198051 | 6.193499 |
| 2 | 0 | 2000-05-06 | 0.323845 | 0.277377 | 0.280454 |
| 3 | 0 | 2000-05-07 | 1.000260 | 1.234950 | 1.244538 |
| 4 | 0 | 2000-05-08 | 2.176284 | 2.252924 | 2.278188 |
evaluate().
| unique_id | ds | y | SeasonalNaive | AutoARIMA | MSTL | |
|---|---|---|---|---|---|---|
| 0 | 0 | 2000-05-04 | 5.267045 | 5.453414 | 5.296218 | 5.283464 |
| 1 | 0 | 2000-05-05 | 6.242415 | 6.136066 | 6.198051 | 6.193499 |
| 2 | 0 | 2000-05-06 | 0.346218 | 0.323845 | 0.277377 | 0.280454 |
| 3 | 0 | 2000-05-07 | 1.134706 | 1.000260 | 1.234950 | 1.244538 |
| 4 | 0 | 2000-05-08 | 2.122063 | 2.176284 | 2.252924 | 2.278188 |
Evaluate models across multiple metrics
evaluate() computes any combination of loss functions from
utilsforecast.losses and returns a tidy DataFrame with one row per
(unique_id, metric) and one column per model.
For Pareto analysis we need one scalar per metric per model — a
single number that summarises performance across all series. The
agg_fn='mean' argument collapses the per-series rows into a single
mean, giving a (n_metrics, n_models) table.
| metric | SeasonalNaive | AutoARIMA | MSTL | |
|---|---|---|---|---|
| 0 | mae | 0.162020 | 0.120070 | 0.119955 |
| 1 | rmse | 0.196461 | 0.145129 | 0.144143 |
| 2 | mape | 0.354416 | 0.359234 | 0.340032 |
| 3 | smape | 0.087060 | 0.065840 | 0.064982 |
Find the Pareto frontier
ParetoFrontier.find_non_dominated() takes the aggregated scores table
and returns only the columns corresponding to non-dominated models —
dropping any model for which another model is at least as good on every
metric and strictly better on at least one.
| metric | MSTL | |
|---|---|---|
| 0 | mae | 0.119955 |
| 1 | rmse | 0.144143 |
| 2 | mape | 0.340032 |
| 3 | smape | 0.064982 |
Focusing on a subset of metrics
You can restrict the comparison to only the metrics that matter for your use case by passing ametrics list.
| metric | MSTL | |
|---|---|---|
| 0 | mae | 0.119955 |
| 1 | rmse | 0.144143 |
Maximization metrics
By default all metrics are minimized (lower is better). If a metric should be maximized — for example, a custom R² score — pass its name inmaximization.
| metric | MSTL | |
|---|---|---|
| 0 | mae | 0.119955 |
| 1 | rmse | 0.144143 |
| 2 | mape | 0.340032 |
| 3 | smape | 0.064982 |
Visualize the 2D Pareto frontier
When comparing two metrics,ParetoFrontier.plot_pareto_2d() renders a
scatter plot where dominated models appear in grey and Pareto-optimal
models appear in red, connected by a dashed frontier line.
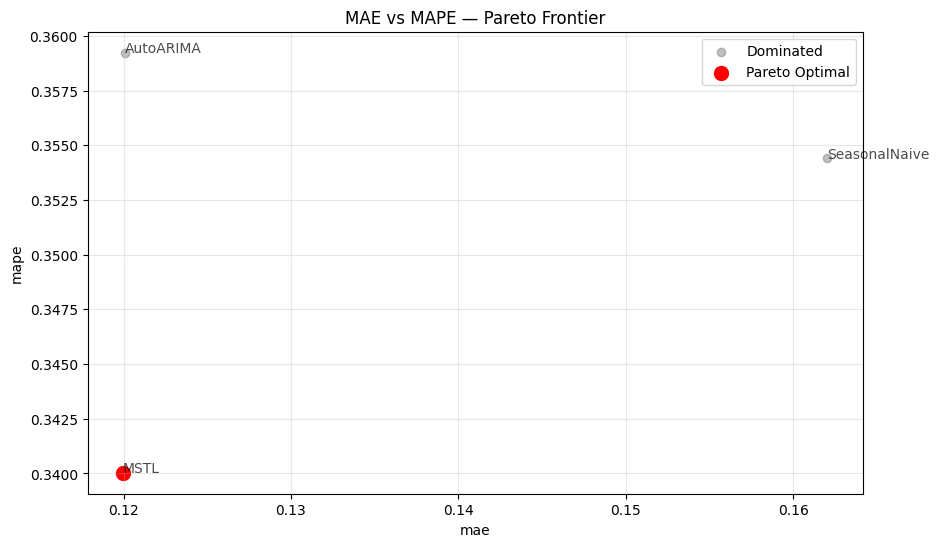
maximize_x and maximize_y flags for metrics where
larger is better, and show_dominated=False to declutter the chart when
many models are present.
Cross-validation: multi-window model selection
A single held-out window can be noisy. StatsForecast’scross_validation() produces estimates across multiple windows, giving
a more robust picture of model performance.
The cross-validation output has a cutoff column — evaluate() keeps
it, so agg_fn='mean' aggregates across series within each cutoff,
not across all windows at once. To collapse everything into a single row
per metric for Pareto analysis, apply a second groupby.
| unique_id | ds | cutoff | y | SeasonalNaive | AutoARIMA | MSTL | |
|---|---|---|---|---|---|---|---|
| 0 | 0 | 2000-05-02 | 2000-05-01 | 3.152391 | 3.061044 | 3.200568 | 3.200882 |
| 1 | 0 | 2000-05-03 | 2000-05-01 | 4.082328 | 4.178149 | 4.194904 | 4.197371 |
| 2 | 0 | 2000-05-04 | 2000-05-01 | 5.267045 | 5.453414 | 5.294926 | 5.284707 |
| 3 | 0 | 2000-05-05 | 2000-05-01 | 6.242415 | 6.136066 | 6.198686 | 6.194711 |
| 4 | 0 | 2000-05-06 | 2000-05-01 | 0.346218 | 0.323845 | 0.277010 | 0.282018 |
| metric | SeasonalNaive | AutoARIMA | MSTL | |
|---|---|---|---|---|
| 0 | mae | 0.163304 | 0.119660 | 0.119076 |
| 1 | rmse | 0.196949 | 0.144753 | 0.143648 |
| 2 | mape | 0.349638 | 0.326070 | 0.317805 |
| 3 | smape | 0.086262 | 0.064798 | 0.063181 |
| metric | MSTL | |
|---|---|---|
| 0 | mae | 0.119076 |
| 1 | rmse | 0.143648 |
| 2 | mape | 0.317805 |
| 3 | smape | 0.063181 |
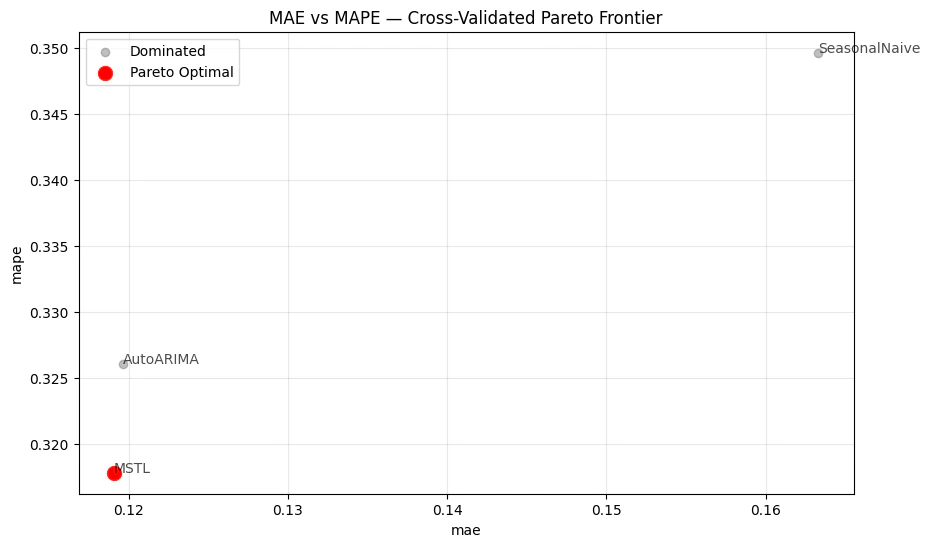
Custom column names
If your pipeline uses column names different from the defaults (unique_id, cutoff), pass id_col and cutoff_col to both
evaluate() and find_non_dominated() so the Pareto analysis correctly
identifies which columns are model predictions.
| metric | MSTL | |
|---|---|---|
| 0 | mae | 0.119955 |
| 1 | rmse | 0.144143 |
Key takeaways
- Single-metric selection discards information. When metrics disagree, there is no universally correct answer — only trade-offs worth making explicit.
- Pareto dominance is a lossless filter. Every eliminated model is objectively outperformed; no information about the surviving frontier is lost.
- Always aggregate before calling
find_non_dominated(). Passagg_fn='mean'toevaluate()so the input has exactly one row per metric. For cross-validation output, apply a second groupby over themetriccolumn to collapse across cutoffs as well. plot_pareto_2d()makes the trade-off tangible. Pick any two metrics on the axes to see which models sit on the frontier and which ones are dominated.- Custom column names are supported. Pass
id_colandcutoff_colconsistently acrossevaluate()andfind_non_dominated()when your data uses non-default names.